status_cols <- c("ND" = "#06D6A0", "ED" = "#1A78C2", "D" = "#E69F00")
df_cea <- data.frame(
effect = c(277.25, 277.57, 277.78, 277.83, 277.76) / 12,
cost = c(26000, 27000, 28020, 28440, 29440),
strategy = c("Status Quo", "One-Time", "Every 5yr", "Every 3yr", "Annual"),
group_label = c("Status Quo", "Infrequent", "Infrequent", "Frequent", "Frequent"),
icon_col = c("person", "vial", "vial", "syringe", "syringe"),
status = factor(c("ND", "ND", "ND", "ND", "D"), levels = c("ND", "ED", "D")),
stringsAsFactors = FALSE
)
dummy_icons <- data.frame(
effect = rep(NA_real_, 3), cost = rep(NA_real_, 3),
icon_col = c("person", "vial", "syringe"),
group_label = factor(c("Status Quo", "Infrequent", "Frequent"),
levels = c("Status Quo", "Infrequent", "Frequent")),
stringsAsFactors = FALSE
)
dummy_status <- data.frame(
effect = rep(NA_real_, 2), cost = rep(NA_real_, 2),
status = factor(c("ND", "D"), levels = c("ND", "ED", "D")),
stringsAsFactors = FALSE
)
dummy_ef <- data.frame(effect = NA_real_, cost = NA_real_,
frontier = "Efficient Frontier")
suppressWarnings(
ggplot(df_cea, aes(x = effect, y = cost, icon = icon_col, color = status)) +
geom_line(data = df_cea %>% filter(status == "ND") %>% arrange(effect),
aes(x = effect, y = cost, group = 1),
color = "#06D6A0", linewidth = 1, alpha = 0.6, inherit.aes = FALSE) +
geom_icon_point(size = 2, dpi = 120, show.legend = FALSE) +
geom_icon_point(data = dummy_icons,
aes(x = effect, y = cost, icon = icon_col, color = group_label),
size = 2, dpi = 120, inherit.aes = FALSE, show.legend = TRUE) +
geom_point(data = dummy_status, aes(x = effect, y = cost, fill = status),
shape = 22, size = 0, alpha = 0, inherit.aes = FALSE, show.legend = TRUE) +
geom_point(data = dummy_ef, aes(x = effect, y = cost, fill = frontier),
shape = NA, size = 0, alpha = 0, inherit.aes = FALSE,
show.legend = TRUE, key_glyph = "path") +
geom_label(data = df_cea %>% filter(status != "ND"),
aes(x = effect, y = cost, label = strategy, fill = status),
color = "white", size = 2.5, vjust = -1.5,
label.size = NA, fontface = "bold",
inherit.aes = FALSE, show.legend = FALSE) +
geom_label(data = df_cea %>% filter(status == "ND"),
aes(x = effect, y = cost, label = strategy),
color = "white", fill = "#06D6A0", size = 2.5,
hjust = -0.1, label.size = NA, fontface = "bold",
inherit.aes = FALSE, show.legend = FALSE) +
scale_color_manual(
name = "HIV Screening",
values = c(status_cols, "Status Quo" = "#2C3E50",
"Infrequent" = "#2C3E50", "Frequent" = "#2C3E50"),
breaks = c("Status Quo", "Infrequent", "Frequent"),
guide = guide_legend(order = 1, ncol = 3,
override.aes = list(alpha = 1, size = 4,
color = "#2C3E50", fill = NA, shape = NA))
) +
scale_fill_manual(
name = "Dominance Status",
values = c("Efficient Frontier" = "#06D6A0", "ND" = "#06D6A0", "D" = "#E69F00"),
breaks = c("Efficient Frontier", "ND", "D"),
labels = c("Efficient Frontier" = "Efficient Frontier",
"ND" = "Non-Dominated", "D" = "Dominated"),
guide = guide_legend(order = 2, ncol = 3,
override.aes = list(
shape = c(NA, 22, 22),
fill = c(NA, "#06D6A0", "#E69F00"),
linetype = c("solid", "blank", "blank"),
color = c("#06D6A0", NA, NA),
linewidth = c(1, 0, 0),
size = c(0, 4, 4),
alpha = 1
))
) +
scale_x_continuous(name = "Effectiveness (QALYs)",
labels = scales::number_format(accuracy = 0.01),
expand = expansion(mult = c(0.05, 0.2))) +
scale_y_continuous(name = "Cost (USD)", labels = scales::dollar,
expand = expansion(mult = c(0.05, 0.2))) +
theme_minimal(base_size = 13) +
theme(legend.position = "bottom", legend.box = "vertical",
panel.grid.minor = element_blank()) +
scale_legend_icon(size = 4, which = "HIV Screening") +
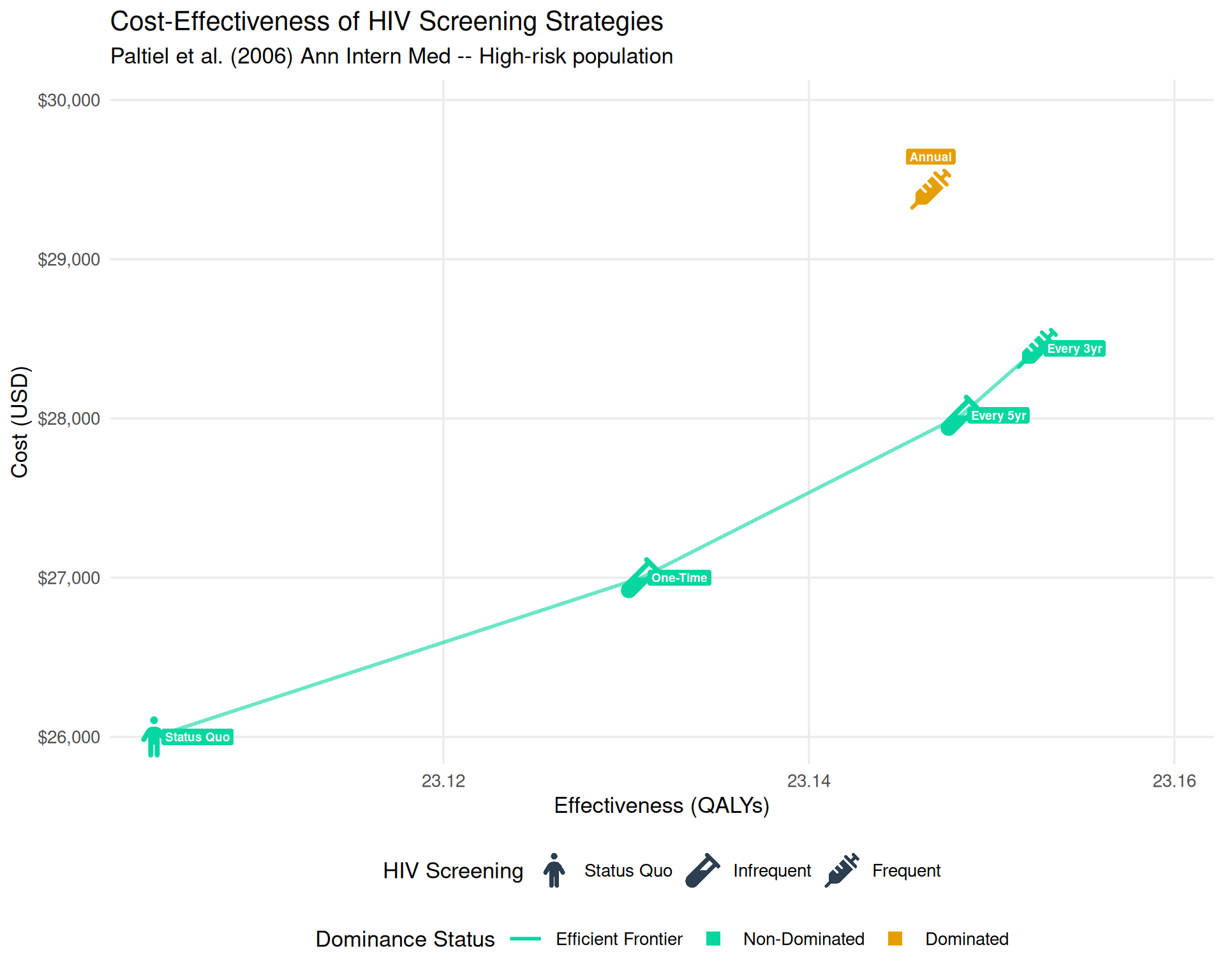
labs(title = "Cost-Effectiveness of HIV Screening Strategies",
subtitle = "Paltiel et al. (2006) Ann Intern Med -- High-risk population")
)